This Fox Chase professor participates in the Undergraduate Summer Research Fellowship.
Learn more about Research Volunteering.
Related Articles
00 / 00

This Fox Chase professor participates in the Undergraduate Summer Research Fellowship.
Learn more about Research Volunteering.
Associate Professor
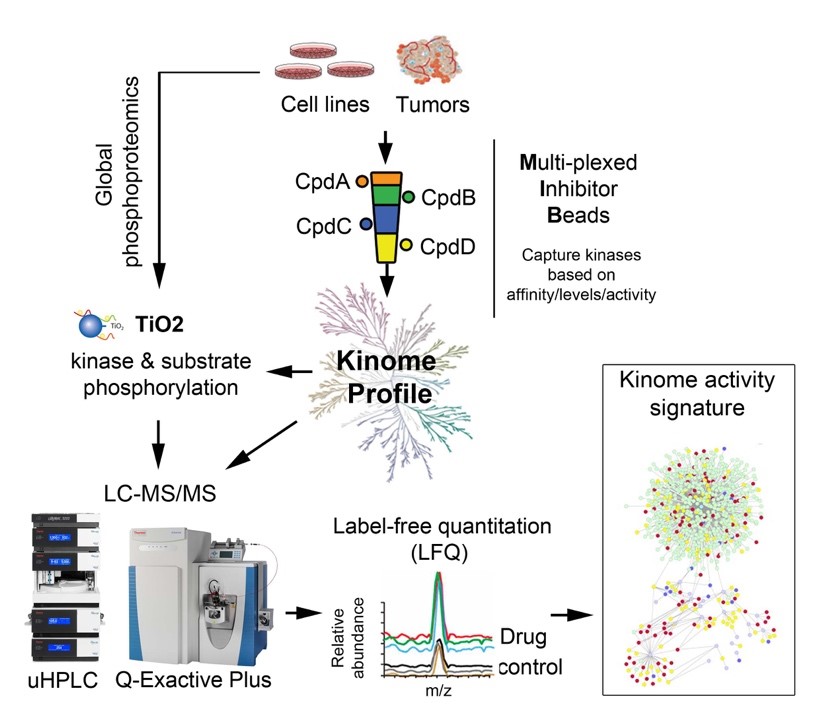
Protein kinases are a family of ~535 enzymes that, collectively, are termed the kinome. Uncontrolled protein kinase activity has been linked to the development of nearly 25% of all cancers; consequently, protein kinases represent one of the most promising avenues for cancer therapy. Despite the high druggability of the kinome, the majority of the kinome remains untargeted, with many kinases having no established oncogenic function. The vast majority of kinase publications focus on a small group of well-understood kinases, yet synthetic lethal screens repeatedly identify various untargeted kinases as playing functional roles in cancer cell proliferation and survival. Our inability to routinely probe these enzymes has hindered previous attempts to understand how they are regulated and function in cancer. Our lab is interested in applying proteomics approaches including kinome profiling and phosphoproteomics to interrogate what is referred to as “the dark cancer kinome” to identify new therapeutic kinase targets for the treatment of cancer.
Protein kinases are a family of ~535 enzymes that, collectively, are termed the kinome. Uncontrolled protein kinase activity has been linked to the development of nearly 25% of all cancers; consequently, protein kinases represent one of the most promising avenues for cancer therapy. Despite the high druggability of the kinome, the majority of the kinome remains untargeted, with many kinases having no established oncogenic function. The vast majority of kinase publications focus on a small group of well-understood kinases, yet synthetic lethal screens repeatedly identify various untargeted kinases as playing functional roles in cancer cell proliferation and survival. Our inability to routinely probe these enzymes has hindered previous attempts to understand how they are regulated and function in cancer. Our lab is interested in applying proteomics approaches including kinome profiling and phosphoproteomics to interrogate what is referred to as “the dark cancer kinome” to identify new therapeutic kinase targets for the treatment of cancer.
Project 1). Unlocking the therapeutic potential of the dark cancer kinome using proteomics.
Our lab has developed an optimized Multiplexed Inhibitor Beads and Mass Spectrometry (MIB-MS) platform (Q-MIBs) specifically designed to profile the kinome of human tissues, providing a quantitative assay for measuring nearly 70% of the kinome in patient tumors. Using this newly designed MIB-MS workflow, our lab comprehensively mapped the kinome landscape of high-grade serous ovarian carcinoma (HGSOC) primary and patient-derived xenograft (PDX) tumors (Science Signaling, 2020). Our proteomics studies of HGSOC tumors revealed a tumor signature comprised of several previously unexplored ‘dark’ kinases, including CDC42BPA (MRCKA). Loss-of-function studies targeting MRCKA in HGSOC cells demonstrated that this previously unexplored kinase was essential for cell motility, growth and survival, as well as spheroid formation, suggesting it is a novel drug target for treating HGSOC. In future studies, we aim to carry out preclinical studies in HGSOC tumor xenograft models to test both the efficacy of MRCKA inhibitors to block tumor growth and cause tumor regression, as well to define potential toxicities associated with targeted MRCKA inhibition.
Similarly, we applied Q-MIBs to map the kinome of endometrial carcinoma (EC) tumors and normal endometrial (NE) tissues and identified several kinases overexpressed in EC, including Serine/Arginine-Rich Splicing Factor kinase (SRPK1). Follow-up loss of function analysis led to the discovery of SRPK1 as an essential kinase regulating mRNA splicing and survival of EC cells, nominating it as a plausible drug target for the treatment of EC (Molecular & Cellular Proteomics, 2020). We are actively pursuing SRPK1 inhibitors alone or in combination with other kinase inhibitors in endometrial cancer xenografts and using proteomics approaches to unravel the mechanisms by which SRPK1 promotes survival of endometrial cancer cells.
Project 2) Applying kinome profiling technologies to predict combination therapies in ovarian cancer.
Although targeted kinase inhibitors show great therapeutic promise, they have had only limited success as single agent treatments due to the rapid development of drug resistance. One mechanism by which tumor cells circumvent the inhibitory action of single agent kinase inhibitors is through ‘kinome reprogramming’, a process characterized by system-wide changes in kinase networks (Cell, 2012). Specifically, tumor cells react to inhibitor treatment by sending alternative signals that cause activation of protein kinase-driven survival pathways that ultimately by-pass the intended drug action, allowing the tumor to escape targeted therapies. These findings suggest that inhibition of multiple kinases might be required to overcome these bypass mechanisms to successfully treat cancer. With numerous kinase-specific inhibitors currently in Phase 1-3 clinical trials for different diseases, development of combination therapies for cancer is a highly tractable goal. Our lab is interested in employing kinome profiling technologies to define the kinases that become activated by specific-targeted therapies to rationally design combination therapies that are predicted to provide more durable therapeutic responses.
Recently, epigenetic therapies that impact transcriptional programs have emerged as promising anti-cancer agents, particular for those cancers exhibiting genomic instability, such as high-grade serous ovarian carcinoma (HGSOC). The BET bromodomain protein BRD4 is overexpressed in ~20% of HGSOC and functions as a transcriptional co-activator promoting tumor growth through the enhanced transcription of key oncogenes, such as MYC. Consequently, BET bromodomain inhibitors (BETi) were developed that displace BET bromodomain proteins from transcriptional complexes disrupting gene transcription. Applying our MIB-MS kinome profiling technologies, we showed, for the first time, that targeting epigenetic proteins, such as BRD4, can significantly rewire the kinome in ovarian cancer cells resulting in activation of a diverse network of kinases facilitating drug resistance (Cell Reports, 2016). Our findings suggest that BET inhibitor therapies may have limited success as single agents in cancer due to adaptive kinome reprogramming and will require combination strategies co-targeting kinases and BET bromodomain proteins. Of particular interest, a revolutionary new class of BRD4 drugs has recently been developed that can selectively remove or degrade BRD4 proteins from cancer cells. These PROTACs (Proteolysis-Targeting Chimeras), highjack the cancer cells protein machinery, effectively tricking the cancer cell into degrading BRD4. Currently little is known about the effectiveness of BRD4-PROTACs in HGSOC cells, nor the potential resistance mechanisms ovarian cancer cells will use to overcome these new drugs. We are currently applying proteomics to unravel the potential resistance mechanisms to BRD4 PROTACs, as well as other PROTAC therapies in ovarian cancer cells, as well as other cancers models.
Targeting the MEK-ERK pathway in ovarian cancer represents another promising therapeutic avenue, particularly in NF1-altered epithelial ovarian cancer (EOC). Loss of RAS suppressor protein Neurofibromin 1 (NF1) frequently occurs in EOC leading to activation of RAF-MEK-ERK signaling, supporting MEK inhibitors for treatment of NF1-altered EOC. Our lab performed a detailed proteomics, genomics and functional analysis of MEK inhibition in NF1-deficient EOC cells and showed MEK inhibitors will likely not be effective as single agent therapies due to drug resistance mediated by kinome reprogramming. Moreover, we showed co-targeting BET bromodomain proteins in combination with MEK inhibitors to block reprogramming at the transcriptional level may provide an epigenetic strategy to overcome MEK inhibitor resistance in NF1-deficient EOC. Currently, we are actively exploring BET protein inhibitors in combination with MEK inhibitors, as well as other kinase inhibitors, such as ERBB receptor therapies.
Project 3) Defining K-Ras mutation-specific kinome signatures and vulnerabilities in colorectal cancer.
Another major focus of our lab has been applying proteomics approaches to explore the impact of KRAS activating mutations on the kinome landscape in colorectal cancers (CRC). KRAS is arguably the most important and most studied oncogene in human cancer, yet despite intensive research effort, there is currently no effective therapy to treat K-Ras mutant tumors. Oncogenic mutations in K-Ras “lock” the enzyme in an ON state resulting in uncontrolled cell growth leading to cancer. Mutations in K-Ras occur in about half of colorectal cancers (CRC) and are associated with poor survival. Distinct mutations in KRAS have been identified in CRC, including mutations in exon 2 - producing (G12C, G12D, G12V, and G13D) and in exon 4 (A146T and A146V). Importantly, there are strong clinical data, and emerging basic science data, indicating that the KRAS mutations in exon 2 are not all alike in their cellular properties. Our lab is currently applying MIB-MS and phosphoproteomics approaches to understand the consequence of these different K-Ras mutations on kinase signaling and determined how these distinct mutations promote resistance to emerging molecular targeted therapies, such as MEK inhibitors.
Ye S., Sharipova D., Kozinova M., Klug L., D’Souza J., Belinsky M.G., Johnson K.J., Einarson M.B., Devarajan K., Zhou Y., Litwin S., Heinrich M.C., DeMatteo R., von Mehren M., Duncan J.S., Rink L., Identification of wee1 as a target in combination with avapritinib for gastrointestinal stromal tumor treatment. JCI Insight. 6(2)2021. PMC7934848. https://www.ncbi.nlm.nih.gov/pubmed/33320833.
Ye S, Sharipova D, Kozinova M, Klug LR, D'Souza JW, Belinsky MG, Johnson KJ, Einarson MB, Devarajan K, Zhou Y, Litwin S, Heinrich MC, DeMatteo RP, von Mehren M, Duncan JS, Rink L. Identification of Wee1 as target in combination with avapritinib for Gastrointestinal Stromal Tumor treatment. JCI Insight, 2020. https://doi.org/10.1172/jci.insight.143474.
Kurimchak AM, Kumar V, Herrera-Montávez C, Johnson KJ, Srivastava N, Devarajan K, Peri S, Cai KQ, Mantia-Smaldone GM, Duncan JS. Kinome Profiling of Primary Endometrial Tumors Using Multiplexed Inhibitor Beads and Mass Spectrometry Identifies SRPK1 As Candidate Therapeutic Target. Molecular & Cellular Proteomics: 19:2068, 2020. https://doi.org/10.1074/mcp.RA120.002012.
Kurimchak AM, Herrera-Montavez C, Brown J, Johnson KJ, Sodi V, Srivastava N, Kumar V, Deihimi S, Peri S, O’Brien S, Puri S, Mantia-Smaldone GM, Jain A, Winters RM, Cai, KQ, Connolly DC, Chernoff J, Duncan JS. Functional proteomics interrogation of the kinome identifies MRCKA as a therapeutic target in high-grade serous ovarian carcinoma. Sci. Signal. 2020 Feb; 13 (619). PMID:32071169.
Kurimchak AM, Shelton C, Herrera-Montavez, C, Duncan KE, Chernoff J, Duncan JS. Intrinsic Resistance to MEK Inhibition Through BET Protein Mediated Kinome Reprogramming in NF1-deficient Ovarian Cancer. Mol Cancer Res. 2019 May 1, doi: 10.1158/1541-7786.MCR-18-1332. PMID: 31043489. PubMed Central PMCID: PMC6679760. Available on 08-01-2020.
Kurimchak AM, Shelton C, Duncan KE, Johnson KJ, Brown J, O'Brien S, Gabbasov R, Fink LS, Li Y, Lounsbury N, Abou-Gharbia M, Childers WE, Connolly DC, Chernoff J, Peterson JR, Duncan JS. Resistance to BET Bromodomain Inhibitors Is Mediated by Kinome Reprogramming in Ovarian Cancer. Cell Rep. 2016 Aug 2;16(5):1273-86. PubMed PMID: 27452461; PubMed Central PMCID: PMC4972668.
Duncan JS, Whittle MC, Nakamura K, Abell AN, Midland AA, Zawistowski JS, Johnson NL, Granger DA, Jordan NV, Darr DB, Usary J, Kuan PF, Smalley DM, Major B, He X, Hoadley KA, Zhou B, Sharpless NE, Perou CM, Kim WY, Gomez SM, Chen X, Jin J, Frye SV, Earp HS, Graves LM, Johnson GL. Dynamic reprogramming of the kinome in response to targeted MEK inhibition in triple-negative breast cancer. Cell. 2012 Apr 13;149(2):307-21. PubMed PMID: 22500798; PubMed Central PMCID: PMC3328787.
This Fox Chase professor participates in the Undergraduate Summer Research Fellowship.
Learn more about Research Volunteering.