Breadcrumb
- Home
- Amy Whitaker
Amy Whitaker, PhD

Educational Background
- PhD, Biochemistry and Biophysics, Texas A&M University, College Station, TX, 2015
- BS, Chemistry, University of St. Thomas, Houston, TX, 2008
Memberships
- American Chemistry Society (ACS), 2023 - Present
- American Society for Biochemistry and Molecular Biology, 2022-Present (ASBMB)
- American Cancer Society Cancer Action Network (ACSCAN), 2018 - Present
- Environmental Mutagenesis and Genomics Society (EMGS), 2016 - Present
- American Association for the Advancement of Science (AAAS), 2015 - Present
Honors & Awards
- EMGS Newly Independent Investigator Engagement Program Awardee, 2022
- 2nd Place Oral Presentation by a New Investigator, EMGS 52nd Annual Meeting, 2021
- Karen and Kelly Gregg Trainee Award, 2021
- NIH Pathway to Independence Award (K99/R00), 2020
- American Cancer Society Postdoctoral Fellowship, 2019
- 1st Place Oral Presentation by a New Investigator, EMGS 50th Annual Meeting, 2019
- 1st Place Oral Presentation by a New Investigator, EMGS 48th Annual Meeting, 2017
- 2nd Place Oral Presentation at Resident, Postdoc, and Fellow (RPF) Research Forum, 2017
People
Additional Staff
Alumni

Haley Kravitz
B.S. Scientific Technician I
Current position: Ph.D. Student in Biochemistry of Health and Disease at Drexel University School of Medicine.

Oviyanna Umoh
Empower Research Fellow; Summer 2023
Current position: Ovi is currently at University of Delaware pursuing her bachelor’s degree in Neurobiology.
Mahi Patel
HS Student Volunteer: Current position: Mahi is currently pursuing her bachelor’s degree in Pure Mathematics at University of Pittsburgh.
Research Interests
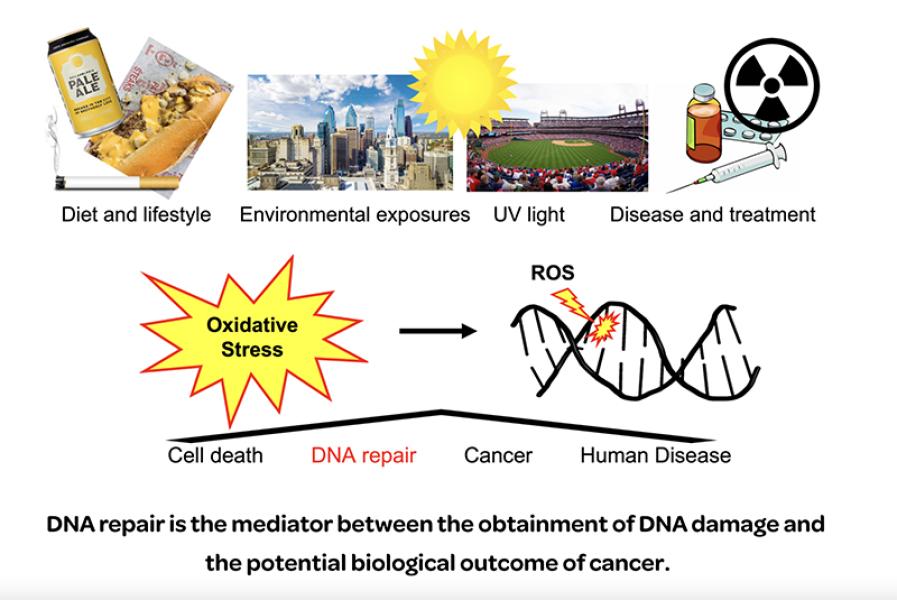
- The epigenetic-like role of oxidative DNA lesions as transcriptional regulators of cancer-associated genes through DNA repair
- DNA repair strategies used to complete oxidative lesion repair in G-quadruplex forming DNA sequences
- DNA structural dynamics in response to DNA damage and repair in G-quadruplexes and other non-canonical DNA secondary structures
Lab Overview
Projects in the Whitaker lab aim to elucidate the molecular mechanisms coupling two key biological responses to oxidative DNA damage within G-quadruplex forming sequences. Specifically, base excision repair (BER) and the epigenic-like transcriptional regulation of key tumor suppressors and oncogenes. The long-term goal is to identify molecular targets that can be exploited to improve cancer therapies.
The primary techniques utilized in the lab include structural biology (X-ray crystallography, Cryo-EM), nucleic acid enzymology, single-molecule fluorescence microscopy, as well as human cell-based assays.
Selected Publications
Deposited and published 32 X-ray crystal structures in the PDB (Protein Data Bank).
DAdil S. Hussen, Haley L. Kravitz, Bret D. Freudenthal, Amy M. Whitaker. Oxidative DNA damage on the VEGF G-quadruplex forming promoter is repaired via long-patch BER. Environ Mol Mutagen. 2023 Aug 22;. PMID: 37606505.
DN. Freund, A. I. Taylor, S. Arangundy-Franklin, N. Subramanian, S. Y. Peak-Chew, Amy M. Whitaker, Bret D. Freudenthal, M. Abramov, P. Herdewijn, Phil Holliger. A two-residue nascent-strand steric gate controls synthesis of 2'-O-methyl- and 2'-O-(2-methoxyethyl)-RNA. Nat Chem. 2023 Jan;15(1):91-100. PMID: 36229679.
Amy M. Whitaker, Wesley J. Stark, Bret D. Freudenthal. Processing oxidatively damaged bases at DNA strand breaks by APE1. Nucleic Acids Res. 2022 Sep 9;50(16):9521-9533. PMID: 36018803.
Max S. Fairlamb, Amy M. Whitaker, Fletcher E. Bain, Maria Spies, Bret D. Freudenthal. 2021. Construction of a Three-Color Prism-Based TIRF Microscope to study the Interactions and Dynamics of Macromolecules. Biology (Basel). PMID: 34201434.
Amy M. Whitaker, Bret D. Freudenthal. 2020. History of DNA polymerase β X-ray crystallography. DNA Repair (Amst). PMID: 33087265.
Amy M. Whitaker, Wesley J. Stark, Tony S. Flynn, Bret D. Freudenthal. 2020. Molecular and structural characterization of disease-associated APE1 polymorphisms. DNA Repair (Amst). Jul-Aug:91-92. PMID: 32454397.
R. McNeill, Amy M. Whitaker, Wesley J. Stark, Jennifer L. Illuzzi, Peter J. McKinnon, Bret D. Freudenthal, David M. Wilson III. 2020. Functions of APE1 that are critical to cell viability and genotoxin resistance. Mutagenesis. 35:27-38. PMID: 31816044. *This paper was awarded paper of the year by Mutagenesis
Amy M. Whitaker, Rockann E. Mosser, Mandar, T. Naik, and Gregory D. Reinhart. 2019. Propagation of the Allosteric Signal in Bacillus stearothermophilus Phosphofructokinase Examined by Methyl-TROSY NMR. Biochemistry. PMID: 31478644.
Amy M. Whitaker, Bret D. Freudenthal. 2018. APE1: A skilled nucleic acid surgeon. DNA Repair (Amst). 71:93-100. PMID: 30170830.
Max S. Fairlamb*, Amy M. Whitaker*, Bret D. Freudenthal. 2018. Apurinic/apyrimidinic (AP) endonuclease 1 processing of AP sites with 5' mismatches. Acta Crystallogr D Struct Biol. 74(Pt 8):760-768. PMID: 30082511. *Co-first authors
Amy M. Whitaker, Tony S. Flynn, Bret D. Freudenthal. 2018. Molecular snapshots of APE1 proofreading mismatches and removing DNA damage. Nature Communications. 9:399. PMID: 9374164.
Amy M. Whitaker*, Mallory S. Smith*, Matthew A. Schaich, Bret D. Freudenthal. 2017. Capturing a mammalian DNA polymerase extending from an oxidized nucleotide. Nucleic Acids Res. PMID: 28449123. *Co-first Authors
Amy M. Whitaker*, Matthew A. Schaich*, Mallory S. Smith, Tony S. Flynn, Bret D. Freudenthal. 2017. Base excision repair of oxidative DNA damage: from mechanism to disease. Front Biosci (Landmark Ed). 22:1493-1522. PMID: 28199214. *Co-first authors
Amy M. Whitaker, Gregory D. Reinhart. 2016. The effect of introducing small cavities on the allosteric inhibition of phosphofructokinase from Bacillus stearothermophilus. Arch Biochem Biophys. 607:1-6. PMID: 27477958.
Additional Publications
Open Positions
ABOUT THE POSITION
A postdoctoral position is available starting immediately for a highly motivated, enthusiastic individual in the NIH-funded group of Dr. Amy Whitaker in the Molecular Therapeutics Program, Fox Chase Cancer Center. The Whitaker group focuses on investigating pressing questions at the intersection of DNA damage, DNA repair, and transcriptional regulation. Potential projects aim to elucidate the mechanistic details by which the repair of oxidative DNA damage within gene regulatory regions of the genome modulate the transcription of key tumor suppressor and oncogenes. The long-term goal is to identify molecular targets that can be exploited to improve cancer therapies. The primary techniques utilized will include structural biology (X-ray crystallography, Cryo-EM), nucleic acid enzymology, single-molecule fluorescence microscopy, as well as human cell-based assays. For more information, please see our website at http://www.dnamywhitakerlab.com/.
Interested applicants should have experience publishing peer reviewed manuscripts and have expertise in either structural biology, single-molecule fluorescence, enzyme kinetics, cancer biology, tissue culture, and/or biochemistry. In addition, applicants should possess excellent written and verbal communication skills, and the ability to conduct research both independently and collaboratively. Candidates must possess a doctoral level degree in a relevant field. Fox Chase postdoctoral fellows enjoy a generous benefits package and salary for this position is set by NIH guidelines and commensurate with experience. Though the position is fully funded, the successful candidate will be mentored and encouraged to apply for additional financial support.
ABOUT THE TRAINING ENVIRONMENT
As one of the four original cancer centers to receive comprehensive designation from the National Cancer Institute, Fox Chase Cancer Center has been at the forefront of cancer research for almost 90 years. We are home to excellent research facilities, top clinicians and scientists, and outstanding patient care. Our singular focus on cancer, which couples discovery science with state of the art clinical care and population health, remains the foundation of our work.
The scientist training programs at Fox Chase Cancer Center provide professional development opportunities in four core areas identified as crucial for successful careers in science, research, and health care including communication, leadership, teaching, and mentorship. Upon joining the program, graduate students and postdocs develop individual development plans to help guide their growth. Training throughout the year is supplemented with free professional development opportunities, including a robust ‘How To’ series, writing courses, networking, mentorship, and teaching opportunities, a trainee-led seminar series, a trainee-led annual Research Conference, and more. Postdocs at Fox Chase Cancer Center are supported by the Temple University Postdoc Association and the Office of Academic Affairs at Fox Chase, and are compensated with competitive pay and benefits.
In addition to the robust training program, scientists at Fox Chase Cancer Center benefit from being part of the rich scientific and biotech environment in the Philadelphia region. Many of our former trainees are now employees (and contacts) at nearby institutions and companies, including The Wistar Institute, Merck, GSK, AACR, and numerous others.
TO APPLY
To apply, please email a CV, a cover letter briefly describing research interest(s) and goals, and the name of at least three references to Dr. Amy Whitaker ([email protected]).






