This Fox Chase professor participates in the Undergraduate Summer Research Fellowship.
Learn more about Research Volunteering.
Related Articles
00 / 00
-

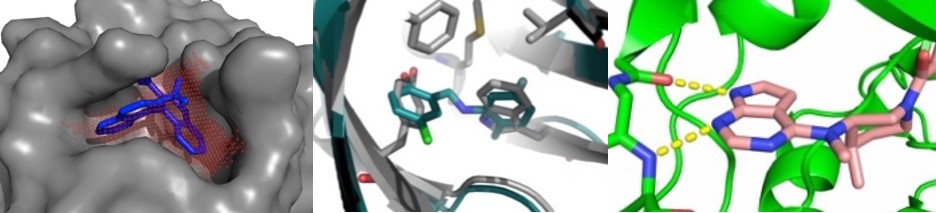
Karanicolas 1200 x 628 px
-

Margie Clapper, co-leader of the Cancer Prevention and Control research program at Fox Chase
-

-

-

-

-

John Karanicolas, PhD


















Patient comments