
Trainee Spotlight: Ting Zhang, PhD
-
Ting Zhang, PhD
Postdoctoral Associate
Dr. Siddharth Balachandran’s Lab
Fox Chase Cancer Center
[email protected]Biography
My journey toward a career in science started in early childhood. I was born and raised in Chongqing the biggest city in southwest China known for its hot summers. As a kid, I was always fascinated by all kinds of small creatures; I desired more than anything to have a dog. In school, I was drawn to biology where I could discuss the natural world with the professor and learn about the origins of the human race.
After graduating from high school, I decided to explore my interests in science in college; therefore, I applied to biological sciences program in Xinan University in Chongqing, China. As part of my senior thesis research, I worked in a Zebrafish lab where I was introduced to animal models and animal development. Through this work, I learned how to utilize animal models to study the earliest stages of development--embryogenesis. To further my understanding of early development, I decided to pursue a PhD in the lab of Dr. Xianming Mo at Sichuan University in China where I studied cell migration during gastrulation to understand how stem cells moved to the “right” place in the embryo to form new tissues. I was amazed by how much could be learned about human development by studying the same processes in model organisms! Although I loved learning about basic developmental processes, I wanted to pursue more practical applications of science. To this end, I moved to the United States to pursue a postdoctoral fellowship with Dr. Wenhui Hu at Temple University in Philadelphia, Pennsylvania. During this period, I explored the new genomic editing tool CRISPR. I utilized this tool to eliminate HIV genomic DNA, which had integrated into a patient’s genome, to provide a promising HIV therapeutic. To expand my skills and learn more virology and immunology, I next joined Dr. Siddharth Balachandran’s lab at Fox Chase Cancer Center as a postdoctoral fellow in 2017. In the Balachandran lab I currently study another virus that significantly impacts human health—influenza virus (IAV)—and a newly described programmed cell death host response called necroptosis. My project has since focused on understanding how cells infected with IAV trigger necroptosis to prevent the spread of IAV to surrounding cells. This year, our findings were published in the journal Cell. In the future, I’m interested in studying the role of necroptosis in immunotherapy.
Research Overview
Each year Influenza A viruses (IAVs), single-stranded RNA viruses, kill or sicken over 10,000 people in the United States. IAVs primarily cause disease through direct infection of the epithelial cells in the respiratory tract where virus replication leads to significant cell death of alveolar epithelia cells, over-recruitment and activation of neutrophils, and dysfunction of respiratory system, which is fatal.
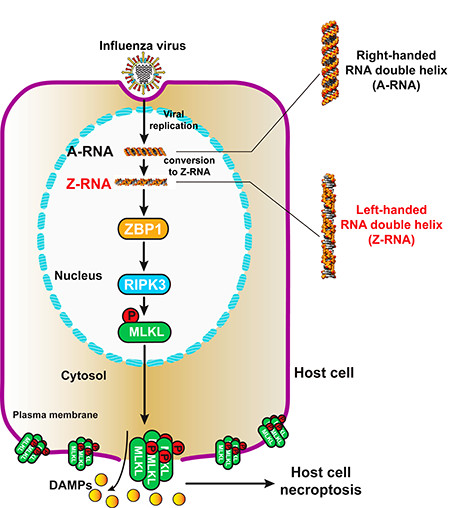
To develop a therapy to block alveolar epithelia cell death and neutrophil over-recruitment induced after IAV infection, we first needed to understand how IAV triggers cell death and activates immune cell recruitment. Previously, our lab uncovered that host protein Z-DNA-binding protein 1 (ZBP1) somehow senses viral RNA, and then activates downstream effectors including receptor interacting serine/threonine kinase 3 (RIPK3) and mixed lineage kinase domain like pseudokinase (MLKL), which lead to a programmed form of necrosis called necroptosis (or inflammatory cell death).
Our current study extends our previous findings and is focused on four questions: first, what viral component does ZBP1 bind; second, where in the host cell does ZBP1 sense this viral ligand; third, how do IAVs cause necroptosis; and fourth, how do IAVs cause pathogenesis? From our previous work, we knew that ZBP1 contained two tandem Zα domains capable of binding to Z-form (left-handed) nucleic acids. To test if ZBP1 binds to the Z-RNA produced by IAVs, we screened all commercial anti-Z-DNA antibodies and found that two of them could recognize Z-RNA. Next, we used these antibodies to analyze the subcellular locational of ZBP1 bound to Z-RNA by immunofluorescence staining. Following IAV infection, we found that ZBP1 sensed Z-RNAs produced by IAV in the nucleus.
To determine how ZBP1-dependent binding to Z-RNAs led to necroptosis, we created ZBP1 ∆Zα mutant which lacks the Z-DNA/Z-RNA binding domain. By co-IP we determined that mutant ZBP1 could no longer induce ZBP1-RIPK3-MLKL complex formation suggesting that ZBP1 binding to viral Z-RNA was critical for activating host necroptosis. We next determined how ZBP1 binding to Z-RNA led to necroptosis. We noticed that ZBP1 and RIPK3 only interact after ZBP1 binds to Z-RNA. From our previous studies and other research articles, we learned that ZBP1 and RIPK3 interact through their respective RHIM domains, which suggested that the RHIM domain of ZBP1 might be exposed after binding to Z-RNA. Then, activated RIPK3 phosphorylates MLKL causing MLKL to aggregate at the nuclear membrane. This aggregation leads to the subsequent rupture of the nuclear membrane and the release or MLKL into the cytoplasm. Once in the cytoplasm, MLKL again accumulated, this time at the cellular membrane, causing rupture of the plasma membrane, and necroptosis.
We next investigated why necroptosis leads to pathogenesis. By ELISA, we observed that plasma membrane rupture of the IAV-infected cell causes the release of damage associated molecular patterns (DAMP) molecules into the extracellular environment. These DAMPs then serve as recruitment signals for immune cells and often the mass over-recruitment of neutrophils to the ruptured cell. Excess neutrophils instigates a damaging feed-forward innate inflammatory circuit which contributes to the pathology induced by IAV. Experimental reduction of neutrophil numbers effectively led to rescue of animals from death in fatal IAV infection.
Together, our findings demonstrate that Z-RNA produced by IAV is a new pathogen-associated molecular pattern, which is recognized by ZBP1 and actives ZBP1-RIPK3-MLKL necroptosis cell death pathway. Necroptosis drives pathogenesis during lethal dose of IAV infections, with potentially important clinical ramifications.
Paper: Zhang et al., 2020, Cell 181, 1115–1129

