This Fox Chase professor participates in the Undergraduate Summer Research Fellowship.
Learn more about Research Volunteering.

This Fox Chase professor participates in the Undergraduate Summer Research Fellowship.
Learn more about Research Volunteering.
Associate Professor
DNA double-strand breaks (DSBs) are among the most deleterious damages to the genome. If unrepaired or improperly repaired, DSBs might lead to gross genomic instability and ultimately many human diseases such as immunodeficiency, premature aging, and, most importantly, cancer. Somewhat paradoxically, cancer cells are also hypersensitive to many chemotherapeutic drugs that act by inducing DSBs. The main focus of our research is to understand the mechanism of DSB repair.
There are four major pathways to repair DSBs in eukaryotes: non-homologous end joining (NHEJ), homologous recombination (HR), single-strand annealing (SSA), and alternative end joining (alt-EJ). The key step in the choice of different DSB repair pathways is the initial processing of DNA ends. NHEJ involves limited processing, but the other three pathways require the resection of 5’ strands to generate 3’ ss-DNA tails. We have found that a major mechanism for resection is carried out by the combined actions of a RecQ-type DNA helicase, the DNA2 ss-DNA nuclease, and the eukaryotic ss-DNA binding protein RPA. The helicase first unwinds the end and DNA2 then degrades the 5’ ss-tail. Both enzymes are stimulated by RPA via physical interactions. Our current research aims to understand how the structure of DNA ends affects the choice of DSB repair pathways and the mechanism of resection and resection-mediated repair.
Kato N., Kawasoe Y., Williams H., Coates E., Roy U., Shi Y., Beese L.S., Scharer O.D., Yan H., Gottesman M.E., Takahashi T.S., Gautier J., Sensing and processing of DNA interstrand crosslinks by the mismatch repair pathway. Cell Rep. 21(5): 1375-1385, 2017. PMC5806701. 7.815
Paudyal S.C., Li S., Yan H., Hunter T., You Z., Dna2 initiates resection at clean DNA double-strand breaks. Nucleic Acids Res. 45(20): 11766-11781, 2017.PMC5714177. 11.147
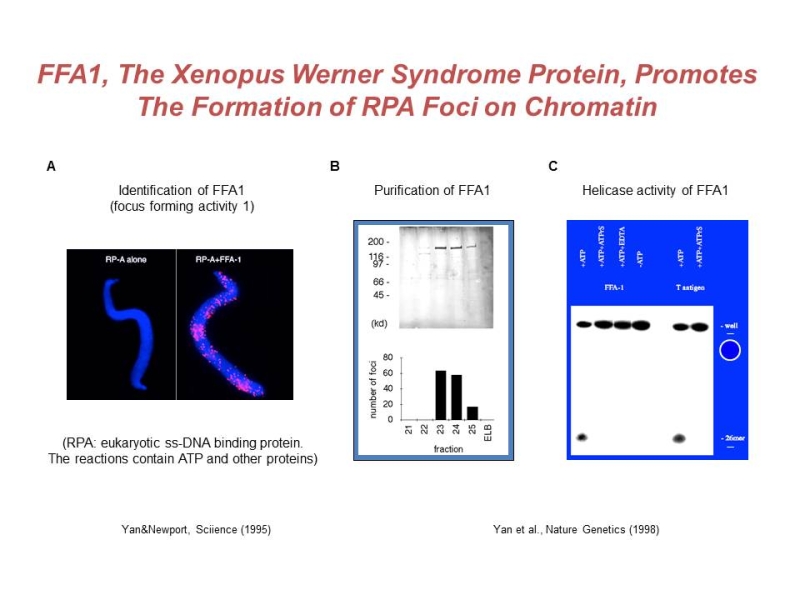
Yan, H., McCane, J., Toczylowski, T., Chen, C. Analysis of the Xenopus Werner syndrome protein in DNA double-strand break repair. J. Cell Biol. 171:217-227, 2005. PMCID: PMC2171202. PubMed
Toczylowski, T., Yan, H. Mechanistic analysis of a DNA end processing pathway mediated by the Xenopus Werner syndrome protein. J. Biol. Chem. 281:33198-33205, 2006. PubMed
Liao, S., Matsumoto, Y., Yan, H. Biochemical reconstitution of abasic DNA lesion replication in Xenopus extracts. Nucleic Acids Res. 35:5422-5429, 2007. PMCID: PMC2018634. PubMed
Liao, S., Toczylowski, T., Yan, H. Identifictaion of the Xenopus DNA2 protein as a major nuclease for the 5’ → 3’ strand-specific processing of DNA ends. Nucleic Acids Res. 36:6091-6100, 2008. PMCID: PMC2577336. PubMed
Yan, H., Toczylowski, T., McCane, J., Chen, C., Liao, S. Replication protein A promotes 5’ → 3’ end processing during homology-dependent DNA double-strand break repair. J. Cell Biol. 192:251-261, 2011. PMCID: PMC3172182. PubMed
Liao, S., Toczylowski, T., Yan, H. Mechanistic analysis of Xenopus EXO1's function in 5'-strand resection at DNA double-strand breaks. Nucleic Acids Res. 39:5967-5977, 2011. PMCID: PMC3152354. PubMed
Liao, S., Guay, C., Toczylowski, T. and Yan, H. Analysis of MRE11's function in the 5'->3' processing of DNA double-strand breaks. Nucleic Acids Res. 40:4496–4506. 2012. PMCID: PMC3378884. PubMed
Tammaro M, Barr P, Ricci B, Yan H. Replication-dependent and transcription-dependent mechanisms of DNA double-strand break induction by the topoisomerase 2-targeting drug Etoposide. PloS one, 8(11):e79202, 2013. PMC3820710. PubMed
Liao, S., Tammaro, M. and Yan, H. (2015) Enriching CRISPR-Cas9 targeted cells by co-targeting the HPRT gene. Nucleic Acids Res. PubMed
Tammaro, M., Liao, S., McCane, J. and Yan, H. (2015) The N-terminus of RPA large subunit and its spatial position are important for the 5'->3' resection of DNA double-strand breaks. Nucleic Acids Res. PubMed
This Fox Chase professor participates in the Undergraduate Summer Research Fellowship.
Learn more about Research Volunteering.